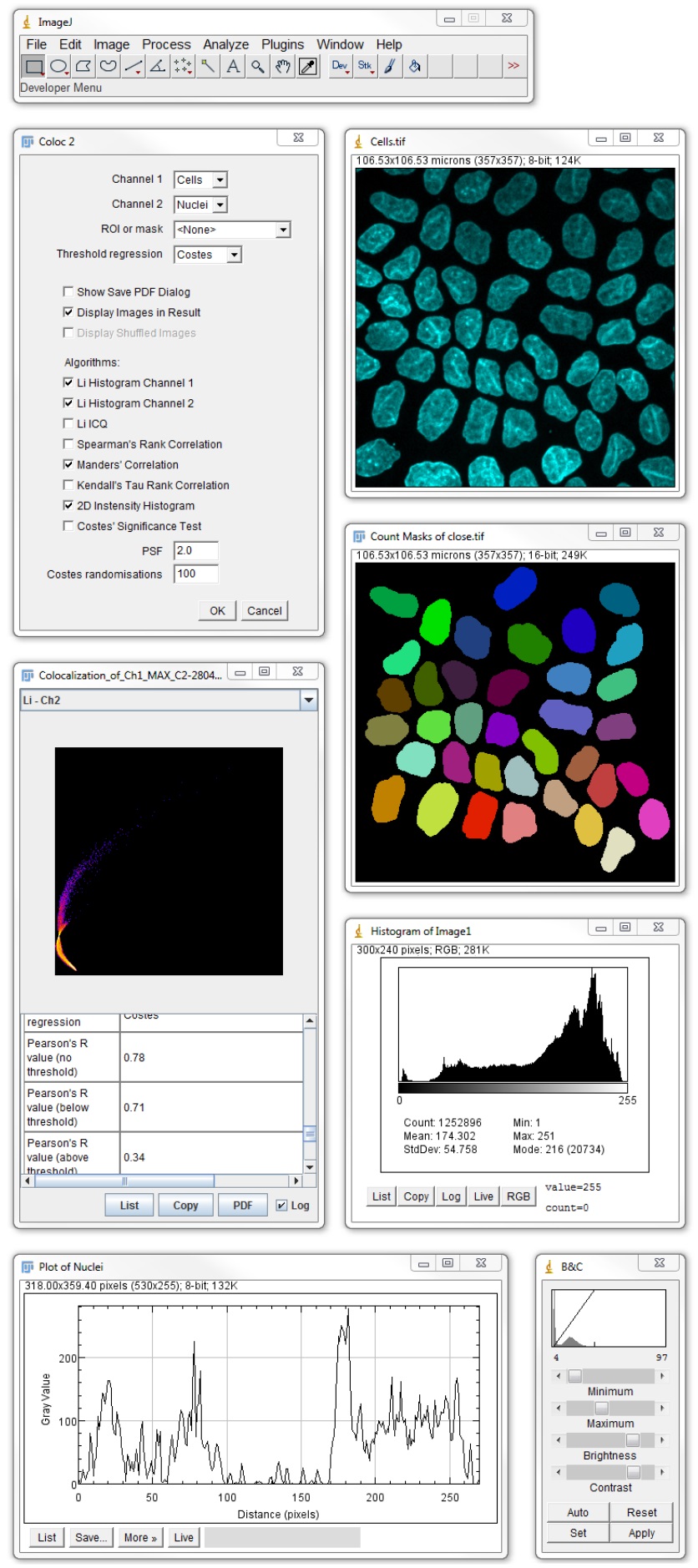
This is particularly relevant to the protooncogene c-fos, the immediate early gene that is directly induced in expression upon neuronal activation, leading to a rapid and transient build-up of c-fos mRNA and consequently of the Fos protein, which decays shortly after cessation of neuronal activity. Signals representing nuanced differences in expression levels, high background and difficult antibodies, in contrast, are not suitable for conventional automated tools. Furthermore, automated tools for the identification of positive cells need high signal-to-noise levels, thus favoring highly expressed proteins. Often, user is required to have image analysis skills and experience, in-depth coding knowledge or access to expensive commercial software. Despite this progress, availability of automated tools that enable easy-to-use, reproducible and reliable identification and quantitative counting of positive cells in immunohistochemical or mRNA in situ hybridization experiments on thick slices of tissue remains limited. A major part of this development has focused on an automated analysis of the expression levels of proteins over regions of interest in histological specimens, such as tumor or immune markers, as clinical diagnostic or prognostic tools. Recently, several automated digital image analysis tools have been developed. Quanty-cFOS facilitates a rapid, accurate and unbiased spatial mapping of neural activity and can also be easily extended to count other types of labelled cells. Here, we demonstrate the application of the tool in a step-by-step manner, with video tutorials, making it easy for novice users to implement. We validated the tool using data from brain sections in response to somatosensory stimuli in a user-interactive manner. This allows for the overcoming of variations in the data and the deriving of cell counts registered to specific brain areas in a highly time-efficient and reliable manner. The algorithms compute the intensity cutoff for positive cells on a user-specified number of images and apply this on all the images to process. Here, we describe a novel open-source ImageJ/Fiji tool, called ‘Quanty-cFOS’, with an easy-to-use, streamlined pipeline for the automated or semi-automated counting of cells positive for the Fos protein and/or c-fos mRNA on images derived from tissue sections.  
Quantitatively analyzing the numbers of cells expressing the Fos protein or c-fos mRNA is a major challenge owing to large human bias, subjectivity and variability in baseline and activity-induced expression. New methods at their own pace.Analysis of neural encoding and plasticity processes frequently relies on studying spatial patterns of activity-induced immediate early genes’ expression, such as c-fos. The classic ImageJ interface, allowing users and developers to migrate to these Maintained such that this new functionality can be seamlessly integrated with Despite the scope of these changes, backwards compatibility is N-dimensional datasets, which are increasingly common in modern imageĪcquisition. The redesigned data model supports arbitrarily large, Its robust new plugin framework allows everythingįrom image formats, to scripting languages, to visualization to be extended by It emphasizes integration with external applications to It separates concerns, fully decoupling the data model from the We present ImageJ2, a total redesign of ImageJ offering a host of newįunctionality. Imaging, ImageJ is at a critical development crossroads. Due to these new and emerging challenges in scientific Imaging paradigms, to ensure the software's ability to handle the requirements User base, diverging plugin suites, and technical limitations have revealed aĬlear need for a concerted software engineering effort to support emerging That spans the biological and physical sciences. Non-programmers, amateur programmers, and professional developers alike.Įnabling such a diversity of contributors has resulted in a large community Due to its ease of use, recordable macro language, andĮxtensible plug-in architecture, ImageJ enjoys contributions from Eliceiri Download PDF Abstract: ImageJ is an image analysis program extensively used in the biological
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
